How Do You Read a Codon Table
Genetic lawmaking
Our genes are encrypted books carrying the secrets of life. In order to understand these secret messages, we would need to know the code and apply the same set of rules, in reverse, to decode it. In this article, we'll have a closer await at the genetic code, which allows DNA and RNA sequences to be "decoded" into the amino acids of proteins.

How do our cells brand proteins – Transcription and Translation
Our genes are written as the nucleotide base of operations pairs (A, T, Chiliad, C) in the DNA. For a gene to exert its function, the genetic information must read out to build a protein. This process is called factor expression.
There are two steps for making proteins from genes:
Starting time, inside the nucleus, a process that makes copies of a sure gene in the form of massager RNAs (mRNAs), called transcription.
Second, these mRNAs are exported outside of the nucleus to the cytoplasm for ribosomes to brand polypeptides/ proteins. This step is called translation.
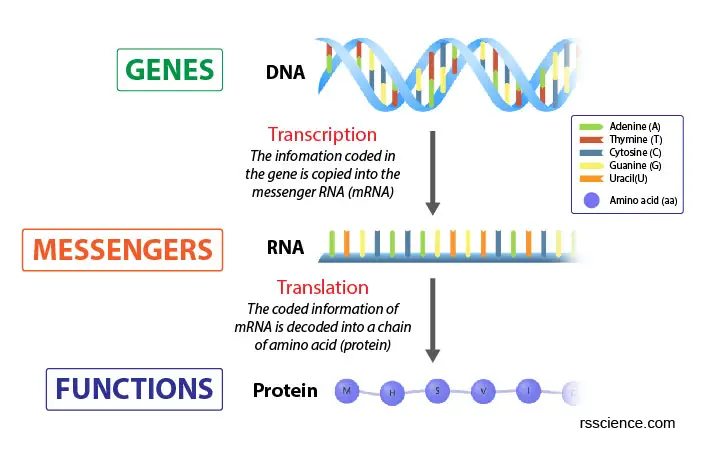
[In this image] The Cardinal Dogma of Biology.
Genes contain the information to build proteins that maintain cell viability. This edifice procedure is washed in ii steps: Transcription and Translation.
The re-create from Deoxyribonucleic acid to RNA is simple: post-obit the complementary base of operations pairing rule. In Deoxyribonucleic acid, there are four nitrogenous base options: Adenine (A), Thymine (T), Cytosine (C), and Guanine (G). Each base can but bail with one other, A with T and C with Thousand. This is called the DNA complementary base of operations pairing rule.
To transcript Dna into mRNA, the dominion is the same. The only difference is that Uracil (U) replaces Thymine (T). And then, G ↔ C, A → U, and T → A. In our cell, the transcription is done by an enzyme chosen RNA polymerase in the nucleus, which can synthesize mRNA from a Deoxyribonucleic acid template.
Dna to mRNA: Using complementary base pairing rules
Knowing this dominion, you can figure out the complementary strand to a single Dna strand based but on the base of operations pair sequence. For example, allow's say y'all know the sequence of i DNA strand that is as follows:
DNA (coding strand): 5'-TTG ACG ACA AGC TGT TTC-three'
Using the complementary base pairing rules, you lot can conclude that the complementary strand is:
DNA (template strand): 3'-AAC TGC TGT TCG ACA AAG-5'
RNA strands are too complimentary with the exception that RNA uses U instead of T. Therefore, y'all can also infer the mRNA strand that would be produced from the first DNA strand. It would be:
mRNA: v'-UUG ACG ACA AGC UGU UUC-three'
RNA to Protein: Using genetic codons
While DNA (genes) and RNA (messengers) use similar codes fabricated of 4 units, proteins are built very differently. Proteins are built using 20 units called amino acids. Translating mRNA to poly peptide becomes much more complicated. To guide this translation, cells follow the genetic code. Co-ordinate to the genetic code, the genetic information is organized in triplets of nucleotides, and each triplet is translated into one amino acid.
For case, the mRNA to a higher place volition translate into
Protein: Leu – Thr – Thr – Ser – Cys – Phe
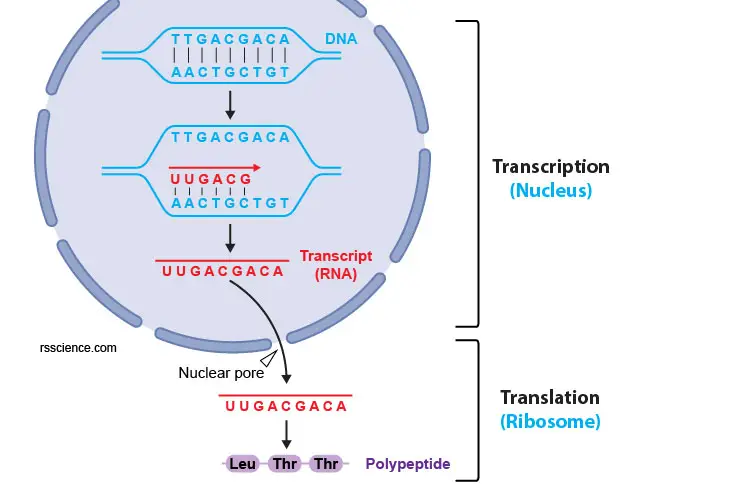
[In this figure] The process of factor expression. From a gene to a protein, there are two steps, transcription and translation. The DNA needs to be transcripted to mRNA using complementary base of operations pairing (i.e., A pairs with U; T pairs with A; C pairs with One thousand; G pairs with C). Next, the mRNAs are exported to the cytoplasm through nuclear pores and are translated to proteins by ribosomes.
Note: A brusque concatenation of amino acids is often referred to every bit a "polypeptide". When the number of amino acids adds up (usually > 30 units) and the polypeptide chain folds into a 3D structure, we call it a "protein".
There are three features of codons:
- Each codon specifies an amino acrid. The full ready of relationships betwixt codons and amino acids is summarized as a Condon Chart or Table.
- 1 "Start" codon (AUG) marks the outset of a protein. AUG encodes the amino acid, called Methionine.
- 3 "Stop" codons mark the end of a protein and terminate the translation.
Who tin can read these codes? Ribosome as a decoding car
Codons in an mRNA are read by a ribosome during translation. A ribosome is a particle-like cell organelle fabricated of ribosomal RNA (rRNA) and ribosomal proteins. A ribosome consists of ii major components: the small and large ribosomal subunits. Three binding sites for tRNA (A, P, and East sites) between the two subunits. Read more nigh ribosomes.
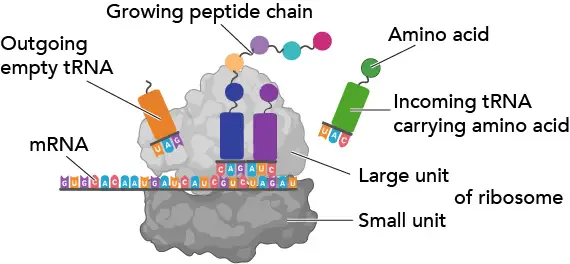
[In this figure] Ribosome.
Ribosomes work like decoding machines to translate the code sequence of mRNA into a protein. Scientists like to phone call ribosomes, the molecular micro-machines, to admire how exquisite the ribosomes' blueprint is!
Transfer RNA (tRNA)
The transfer RNA (tRNA) is one type of RNA molecule. Its job is to carry the amino acid that matches the mRNA codon to the ribosome.
The tRNA contains a three-letter of the alphabet code on one side and carries a specific amino acid on the other side. The code on tRNA (called an anticodon) must match the three-letter code (the codon) on the mRNA already in the ribosome. The particular amino acid that tRNA carries is determined by a 3-letter anticodon it bears. For instance, if the three-letter of the alphabet code is AUG on mRNA, the tRNA that carried Methionine (Met) will be selected and recruited to the ribosome. This is an essential part of the translation process, and it is surprising how few "errors of translation" occur.
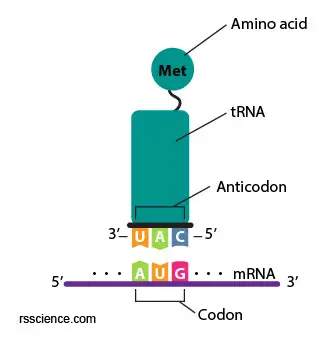
[In this figure]A anticodon UAG on the tRNA matches to the AUG on the mRNA (complimentary) and bring the right amino acrid (Methionine) to the ribosomes.
Protein translation begins with a first codon (always AUG → Methionine) and continues until a stop codon (any one of the iii: UAA, UAG, or UGA) is reached. mRNA codons are read from 5′ end to 3′ end, and its order specifies the gild of amino acids in a protein from N-terminus to C-terminus.
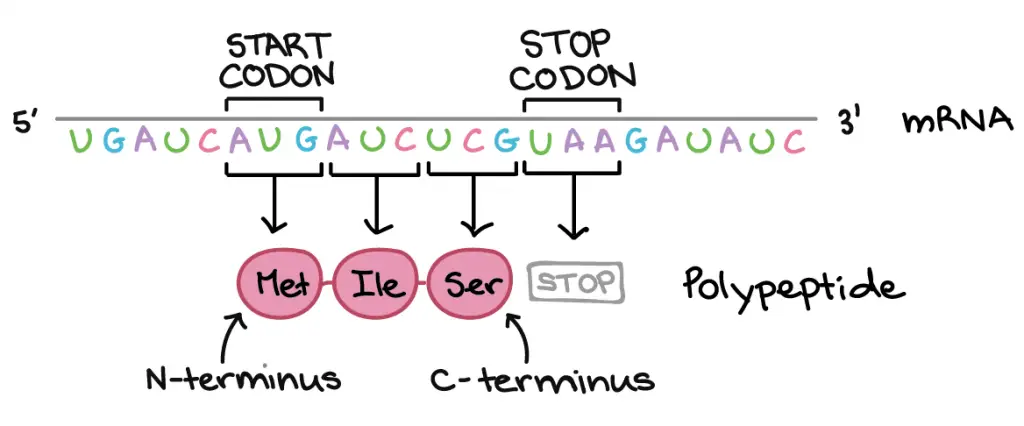
[In this figure]Directionality: Deoxyribonucleic acid and RNA read from 5' end to iii' terminate. Instead, proteins or polypeptides read from Due north-terminus (amino grouping) to C-terminus (carboxyl group). The beginning and the end of a translation is marked past the Start and Finish codons, respectively.
Photo credit: khanacademy
The amino acids codon chart
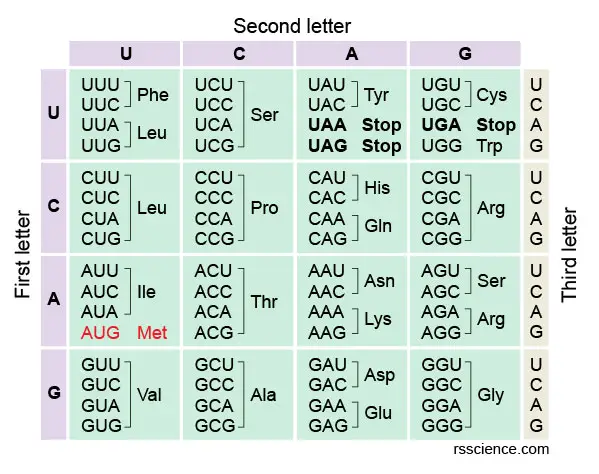
The full prepare of relationships betwixt codons and amino acids (or stop signals) is called the genetic lawmaking. The genetic code is oftentimes summarized in a codon chart (or codon table), where codons are translated to amino acids.
[In this paradigm] Condon ready tin can also be presented as a codon bicycle.
Photo credit: wiki
How do you read the codon chart?
The codon chart may await intimidating at kickoff. In fact, it is not difficult at all once y'all understand its rule.
Let'south take codon ACU as an example. If yous want to know which amino acid ACU encodes, you beginning wait at the left side of the table. Find the "A" on the axis of the left side, which refers to the first alphabetic character of the codon triplet. All these codons starting with "A" are in this row.
Next, nosotros wait at the top of the table. This upper axis indicates the 2nd letter of the alphabet of the codon triplet. In one case we notice "C along the upper axis, it tells us nearly the cavalcade in which our codon will be found. Find the intersecting box of "A" row and "C" cavalcade in the table. You lot will see this box containing four codons and easily find the one you're looking for.
In our example, ACU encodes Thr (or Threonine). You may also find that all ACU, ACC, ACA, and ACG encode the same amino acid. Observe that many amino acids are represented in the table past more than i codon. For instance, there are vi different means to "write" Leucine in the linguistic communication of mRNA (meet if you lot can find all six).
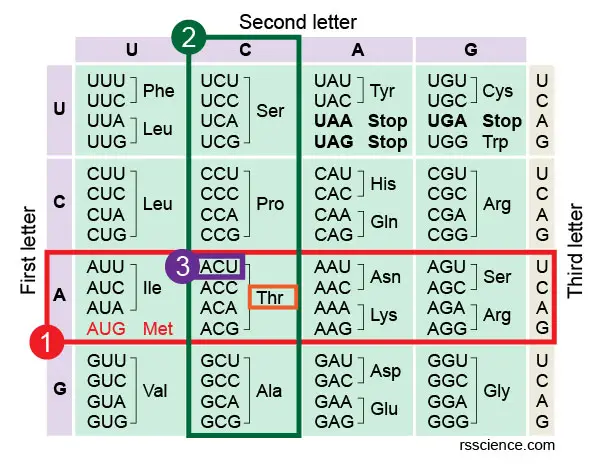
[In this image] How to read the amino acids codon chart?
Following Step 1-iii to find the codon triplet in the table.
In this table, you lot can besides meet that UAA, UAG, and UGA do not encode any amino acrid, pregnant they are stop codons.
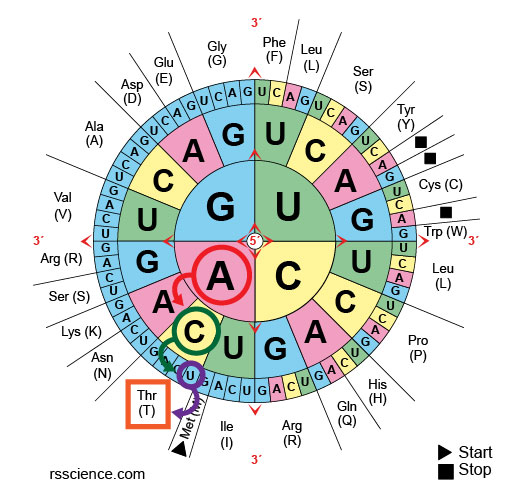
[In this epitome] For a codon wheel, the rule is the same: start from the center to find the first letter of triplet, so move toward the periphery for 2nd and 3rd letters.
You and your family or classroom can play the "Codon Bingo" to get familiar with the genetic code. Here is a downloadable version.
Reference Table: a summary of all amino acids codons
| Amino Acid | Codon |
| Phenylalanine (Phe) | UUU, UUC |
| Leucine (Leu) | UUA, UUG, CUU, CUC, CUA, CUG |
| Methionine (Met) / Beginning Codon | AUG |
| Valine (Val) | GUU, GUC, GUA, GUG |
| Serine (Ser) | UCU, UCC, UCA, UCG, AGU, AGC |
| Proline (Pro) | CCU, CCC, CCA, CCG |
| Threonine (Thr) | ACU, ACC, ACA, ACG |
| Alanine (Ala) | GCU, GCC, GCA, GCG |
| Tyrosine (Tyr) | UAU, UAC |
| Histidine (His) | CAU, CAC |
| Glutamine (Gln) | CAA, CAG |
| Asparagine (Asn) | AAU, AAC |
| Lysine (Lys) | AAA, AAG |
| Aspartic Acid (Asp) | GAU, GAC |
| Glutamic Acid (Glu) | GAA, GAG |
| Cysteine (Cys) | UGU, UGC |
| Tryptophan (Trp) | UGG |
| Arginine (Arg) | CGU, CGC, CGA, CGG, AGA, AGG |
| Glycine (Gly) | GGU, GGC, GGA, GGG |
| Isoleucine (Ile) | AUU, AUC, AUA |
| Stop Codon | UAA, UAG, UGA |
Molecular structures of Amino acids
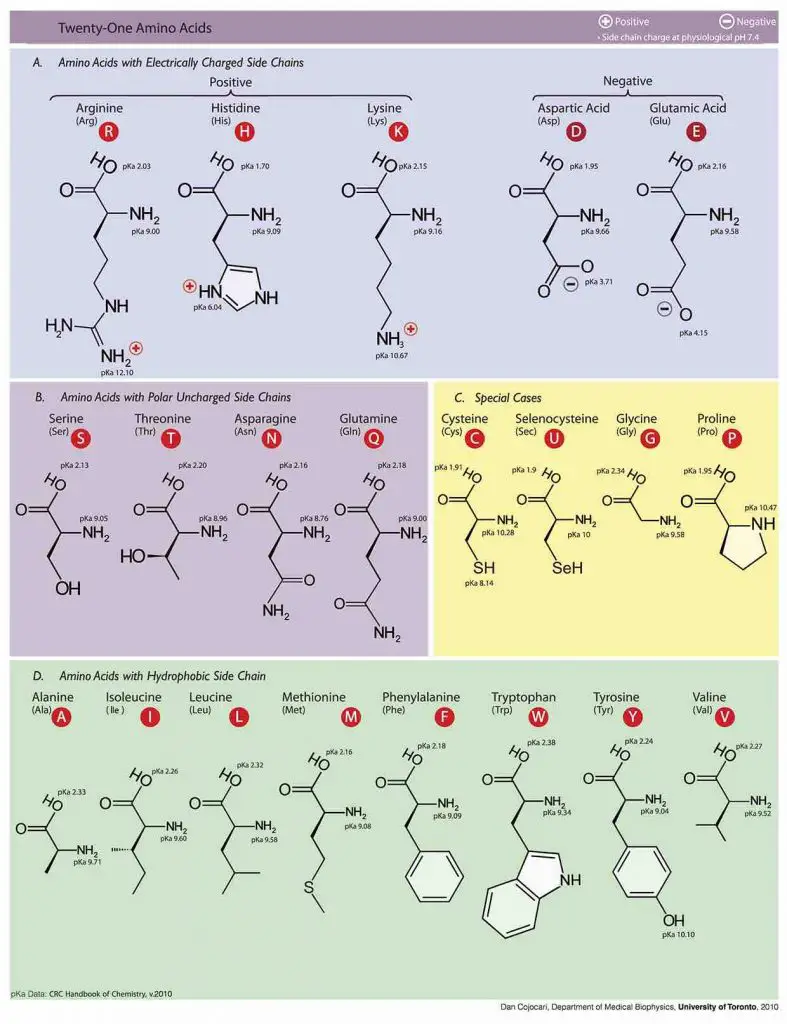
Photo source: wiki
Standard Genetic Code
The genetic code we mentioned here is universal; with only a few exceptions, virtually all species (from bacteria to humans) utilise the same set of standard code. Some ciliates, such as Paramecium bursaria, use unusual genetic lawmaking.
Another exception is mitochondrial DNA. Mitochondria have their ain copies of Deoxyribonucleic acid as well as an independent organisation of ribosomes and tRNAs. If you are non familiar with mitochondria, click here to learn more virtually mitochondria.
The mitochondrial lawmaking is slightly dissimilar from the standard genetic lawmaking. Moreover, dissimilar species take their own versions of mitochondrial codes. For instance, our (vertebrate) mitochondrial code is different from the one yeast uses. AGA and AGG encode Arginine (Arg) in the standard genetic code. However, AGA and AGG act as stop codons in the vertebrate mitochondrial code. In addition, UGA and AUA modify from stop codon and Isoleucine (Ile) to Methionine (Met) and Tryptophan (Trp), respectively, in mitochondria.
The same situation also happens in the plant's chloroplast and plastid codes.
Codon usage biases
Although most organisms use the standard code, nonetheless, they may have their own biases in terms of choosing which codons to use. For instance, baking yeasts prefer using UGU for Cysteine. In contrast, in human cells, nosotros prefer UGC.
Codon usage biases could be the outcome of natural option (tRNA abundance). For laboratories to produce certain proteins in a large quantity, researchers may perform "codon optimization" to resynthesize genes in such a way that their codons are more advisable for the desired expression host (i.e., making human proteins in E coli. bacteria).
What is reading frame?
Since the DNA sequence is read past triplets, starting from which letter (or reading frame) becomes a critical problem.
Permit's look at an instance. The mRNA below can be translated into 3 totally unlike orders of amino acids, depending on the frame in which it'south read. How practise our cells know which of these proteins to brand?
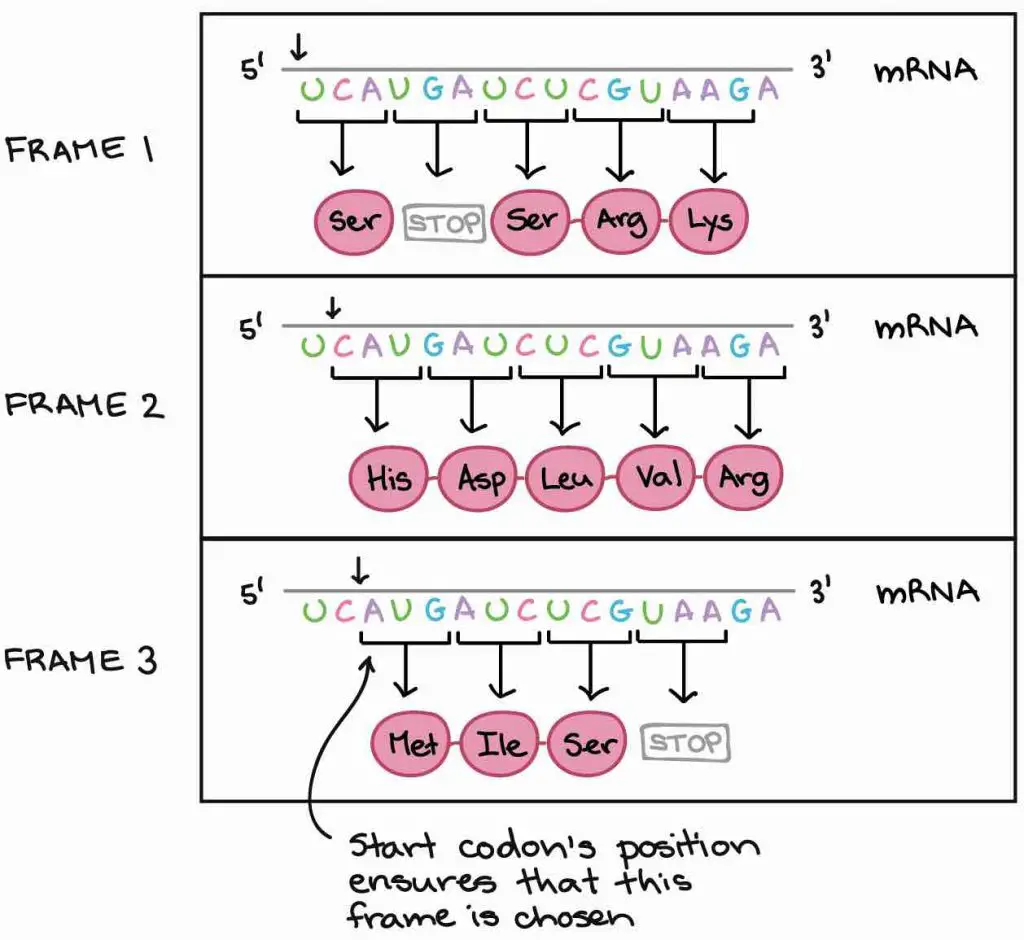
[In this image] Iii possible reading frames could atomic number 82 to totally different results.
Photo credit: khanacademy
Our cells use a very smart strategy to solve this trouble – the "showtime codon". Because the translation only begins at the start codon (AUG) and continues in successive groups of three, the position of the showtime codon ensures that the mRNA is read in the correct frame (in the example above, in Frame 3).
What happens if the Deoxyribonucleic acid sequences are messed up – Mutation
Mutations (changes in DNA sequences) may derail the genetic data and cause cells to make the incorrect proteins. Mutations are the major cause of cancers and many genetic disorders.
Even a single base pair contradistinct (called point mutation) can cause a significant upshot. Bespeak mutations tin have one of iii effects.
Silent mutation
Showtime, the base of operations exchange can be a silent mutation where the altered codon corresponds to the same amino acid. For example, changing from UCU to UCC has no issue since both codons as encode Serine (Ser).
Missense mutation
Second, the base exchange tin can exist a missense mutation where the contradistinct codon corresponds to a dissimilar amino acrid. For example, changing from UCU to UGU will plough Serine (Ser) to Cysteine (Cys). If this mutation happens in the disquisitional region (i.east., enzymatic site) of the protein, a bespeak mutation can mess up the whole poly peptide function.
Nonsense mutation
Third, the base substitution can be a nonsense mutation where the altered codon becomes a stop bespeak. This is the worst cause because the translation will finish too early, resulting in a truncated protein.
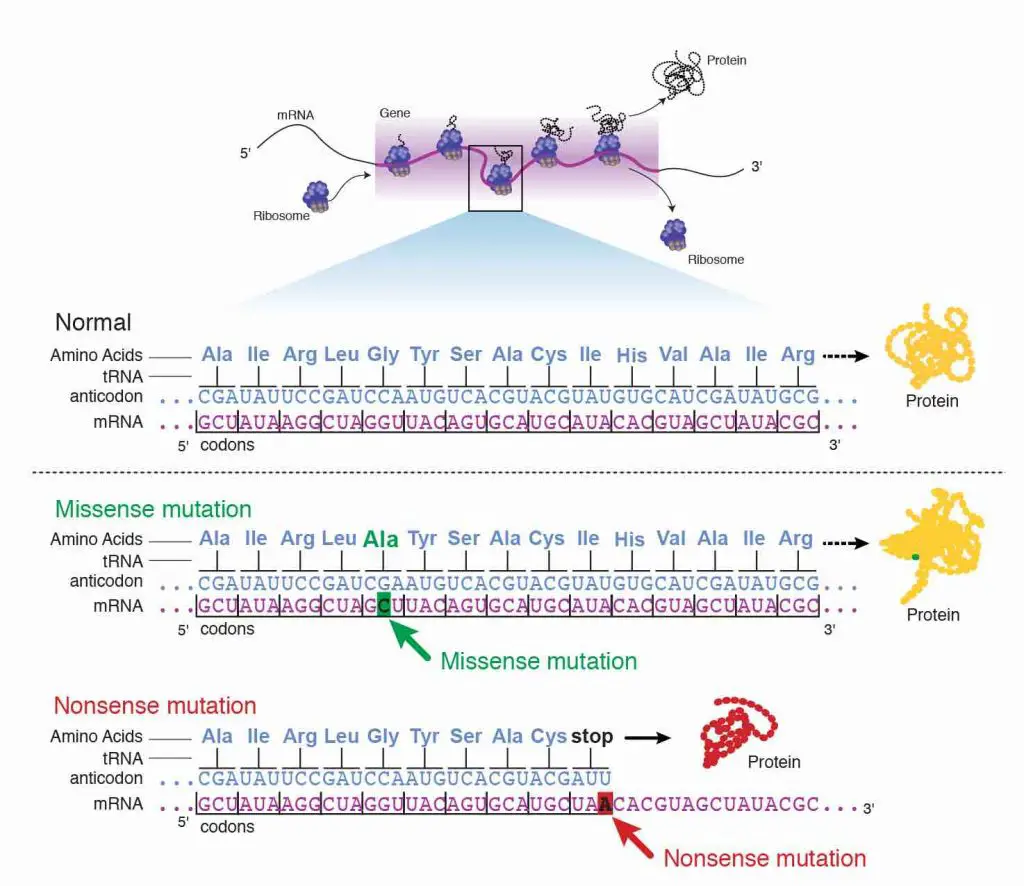
[In this image] The examples of showing the consequence of missense mutation and nonsense mutation.
Photo credit: NIH
Mutations could too happen when nucleotides are inserted or deleted from the original DNA sequence. The insertion or deletion of "one or two" nucleotides can change the reading frame (frameshift mutation). A frameshift tin totally mess up the amino acid orders "downstream" the mutation site.
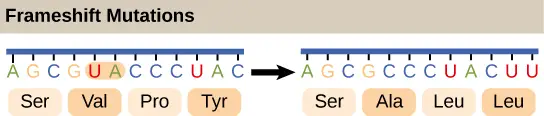
[In this image] The example of showing the effect of frameshift mutation.
Photograph credit: openstax
How was the genetic code discovered?
The understanding of genetic code is the foundation of modem biotechnology. Without the ability to read the Dna information, many exciting techniques and therapies, including personalized medicine, gene therapy, CRISPR gene editing, and recombinant protein drugs, won't exist.
To cleft the genetic code, researchers needed to effigy out how nucleotides sequences in a Dna or RNA molecule could encode the sequence of amino acids. In the mid-1950s, physicist George Gamow predicted that the genetic code is likely composed of triplets of nucleotides – because the possible combination of duplet is non enough (4×4 = 16), and that of quadruplet is too many (4x4x4x4 = 256), to cover 20 kinds of amino acids.
The actual experiments to pinpoint the genetic lawmaking began in 1961 by American biochemist Marshall Nirenberg. Nirenberg was able to link the relationships betwixt nucleotide triplets to particular amino acids by two experimental innovations:
- He can synthesize artificial mRNA molecules with specific, known sequences.
- He had a system to translate mRNAs into polypeptides exterior of a cell (a "cell-gratis" system). Nirenberg did and then in a test tube of cytoplasm from flare-up Due east. coli leaner, which contains all the ingredients needed for translation.
Nirenberg started with an mRNA molecule consisting only of the nucleotide uracil (called poly-U). When he added poly-U mRNA to the jail cell-free organization, he found that the polypeptides made consisted exclusively of the amino acid – Phenylalanine (Phe). Nirenberg concluded that UUU might code for phenylalanine. Using the same approach, he discovered triplet CCC codes for Proline (Pro).

Photo credit: khanacademy
Following this concept, the biochemist Har Gobind Khorana extended Nirenberg's experiment by synthesizing bogus mRNAs with more than complex sequences. By 1965, Nirenberg, Khorana, and their colleagues had deciphered the unabridged genetic code.
For their contributions, Nirenberg and Khorana (forth with another genetic lawmaking researcher, Robert Holley) received the Nobel Prize in Physiology or Medicine in 1968.

[In this paradigm] The Nobel Prize in Physiology or Medicine 1968 was awarded jointly to Robert W. Holley, Har Gobind Khorana, and Marshall W. Nirenberg "for their interpretation of the genetic lawmaking and its function in poly peptide synthesis."
Photo credit: The Nobel Prize
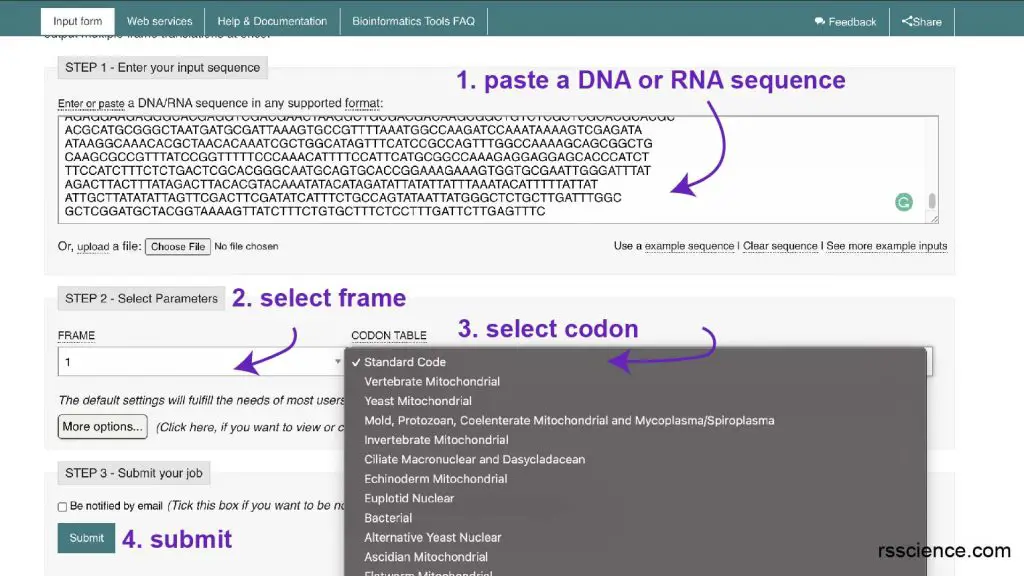
In my research that I demand to clone a particular Deoxyribonucleic acid for poly peptide expression, I typically use EMBOSS Transeq from EMBL-EBI.
Step one: Paste a piece of DNA sequence, yous can apply their example sequence.
Step 2: Select the reading frame you want to utilize
Pace three: Select codon. I typically use standard code.
Step iv: Hit submit
Step 5: Wait for the protein sequence upshot!
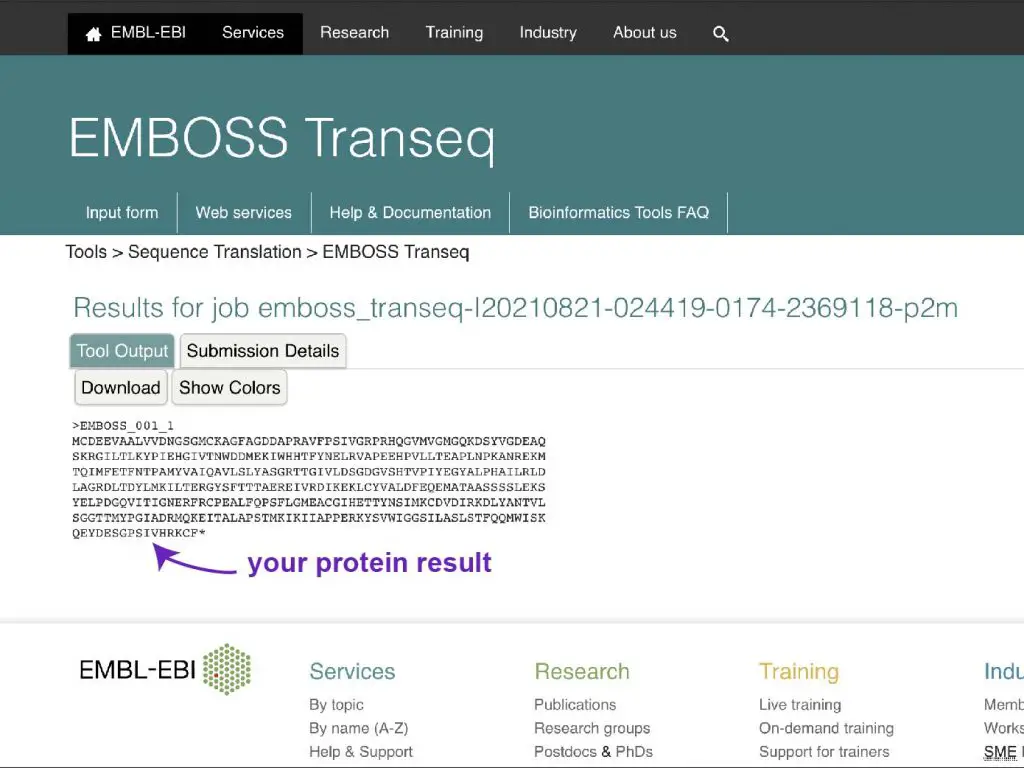
References
landerosbobjecied.blogspot.com
Source: https://rsscience.com/codon-chart/
0 Response to "How Do You Read a Codon Table"
Post a Comment